|
 |
| jbm > Volume 29(3); 2022 > Article |
|
Abstract
Background
Methods
Results
DECLARATIONS
Funding
This work was supported by National Research Foundation of Korea (NRF) grants funded by the Korean government (MSIT) (2019R1A5A2027521).
Fig. 1

Fig. 2
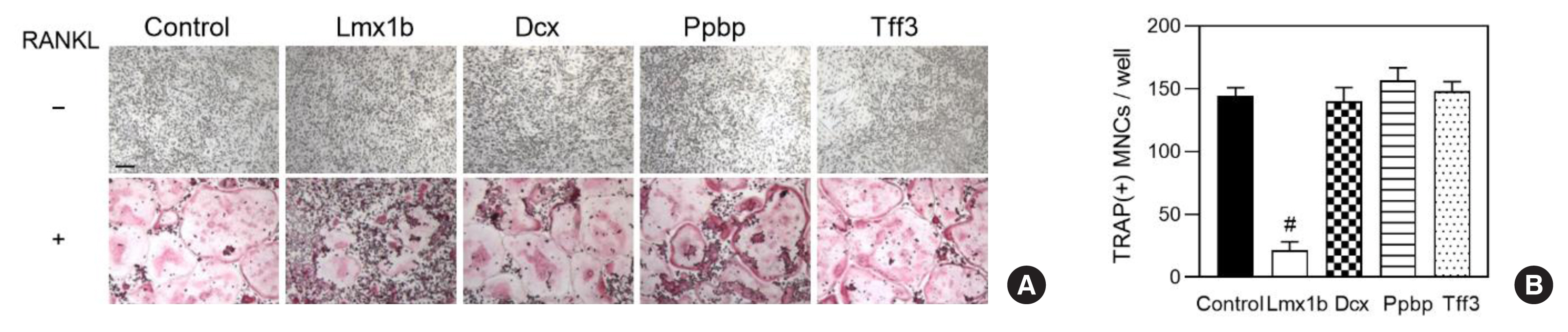
Fig. 3
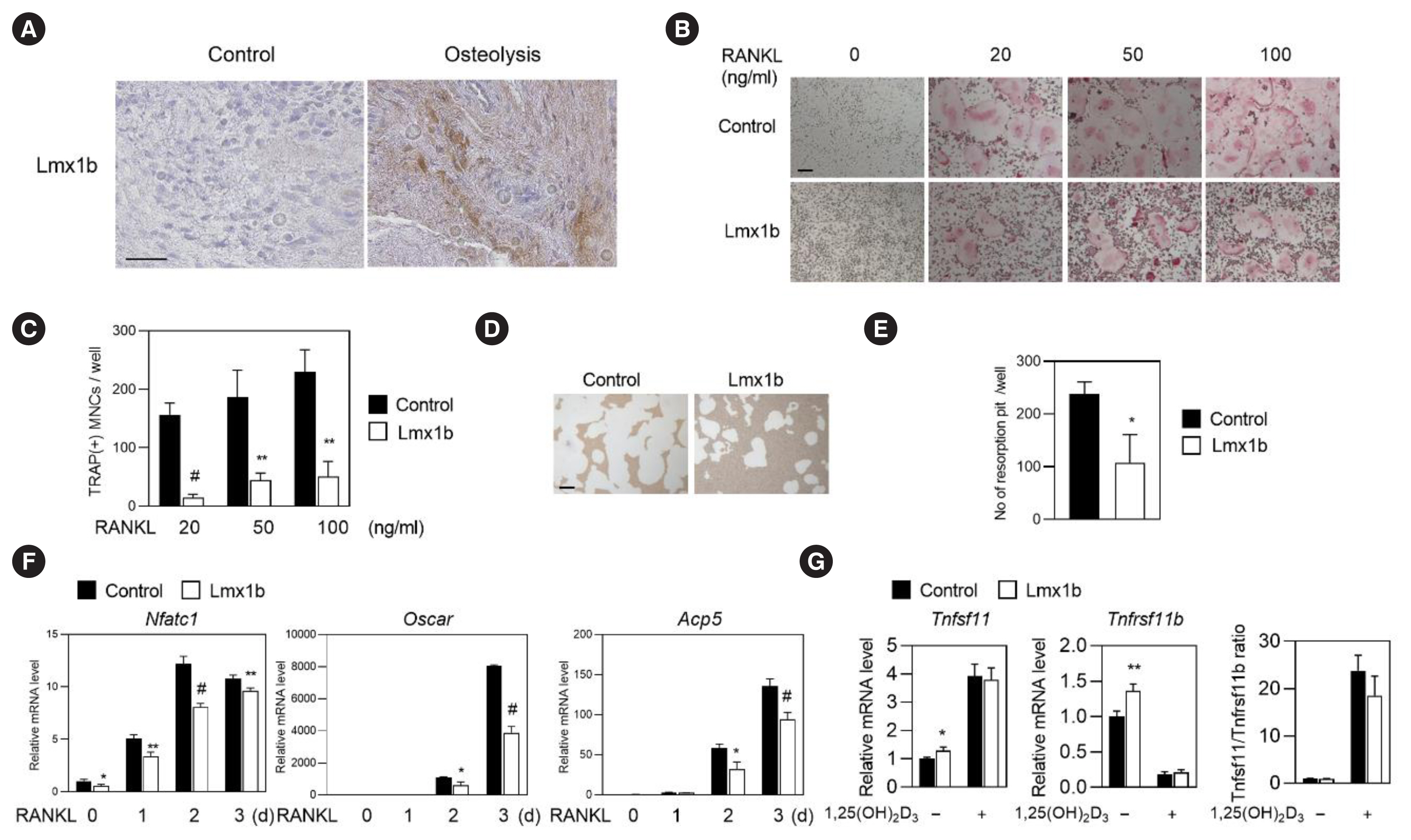
Fig. 4
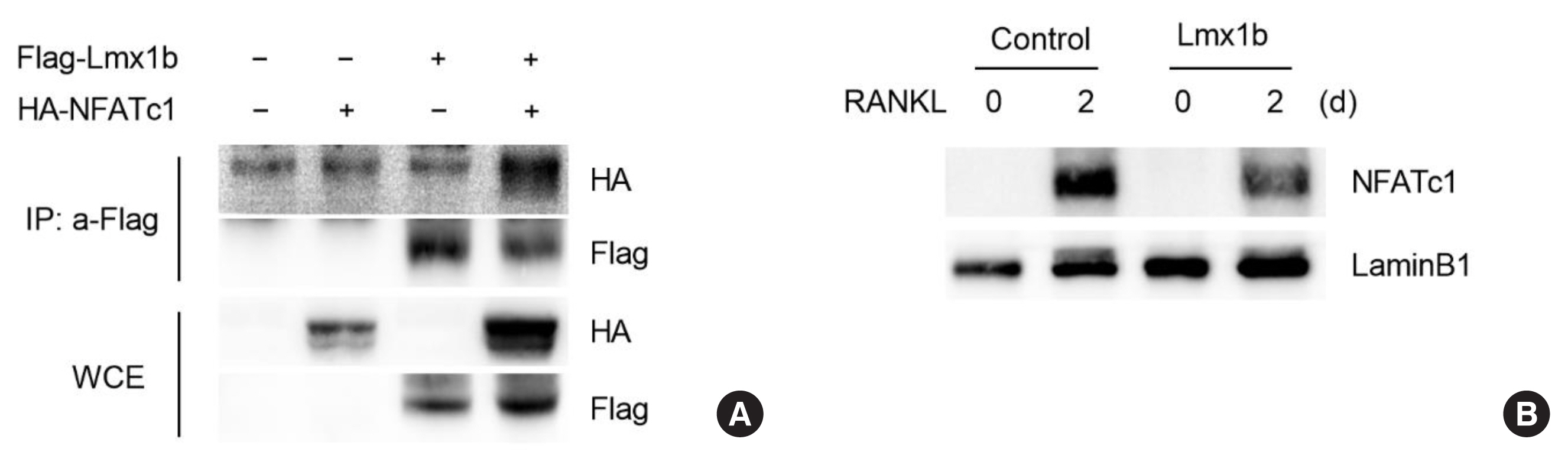
Table 1
Lmx1b, LIM homeobox transcription factor 1-β; PPBP, pro-platelet basic protein; TFF3, trefoil factor 3; DCX, doublecortin; POU2F3, POU class 2 homeobox 3; RNF165, ring finger protein 165; LTB4R2, leukotriene B4 receptor 2; PAK1, p21-activated kinase 1; FERMT1, FERM domain containing kindlin 1; VWA2, von Willebrand factor A domain containing 2; OPRM1, opioid receptor mu 1; TNFSF11, tumor necrosis factor ligand superfamily member 11; TNFRSF11B, tumor necrosis factor receptor superfamily member 11b; TNFRSF11A, tumor necrosis factor receptor superfamily member 11a; GAPDH, glyceraldehyde 3-phosphate dehydrogenase; NFATc1, nuclear factor of activated T cells 1; Oscar, osteoclast-associated receptor; Acp5, acid phosphatase 5.
REFERENCES
- TOOLS
-
METRICS

-
- 6 Crossref
- 0 Scopus
- 2,465 View
- 57 Download
- ORCID iDs
-
Kabsun Kim

https://orcid.org/0000-0002-8320-5030Jeong Eun Han

https://orcid.org/0000-0002-0515-1890Keun-Bae Lee

https://orcid.org/0000-0003-3808-2649Nacksung Kim

https://orcid.org/0000-0002-8132-3981 - Related articles





 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print